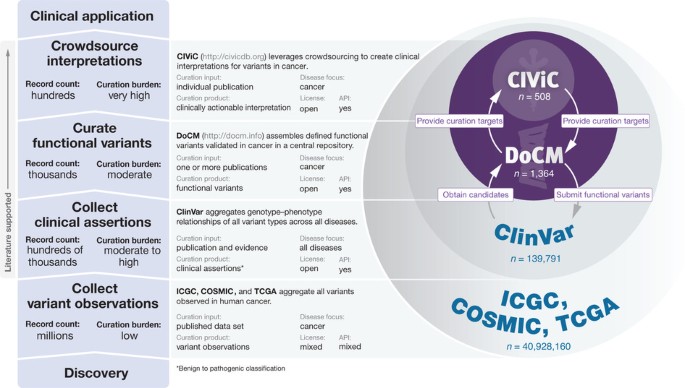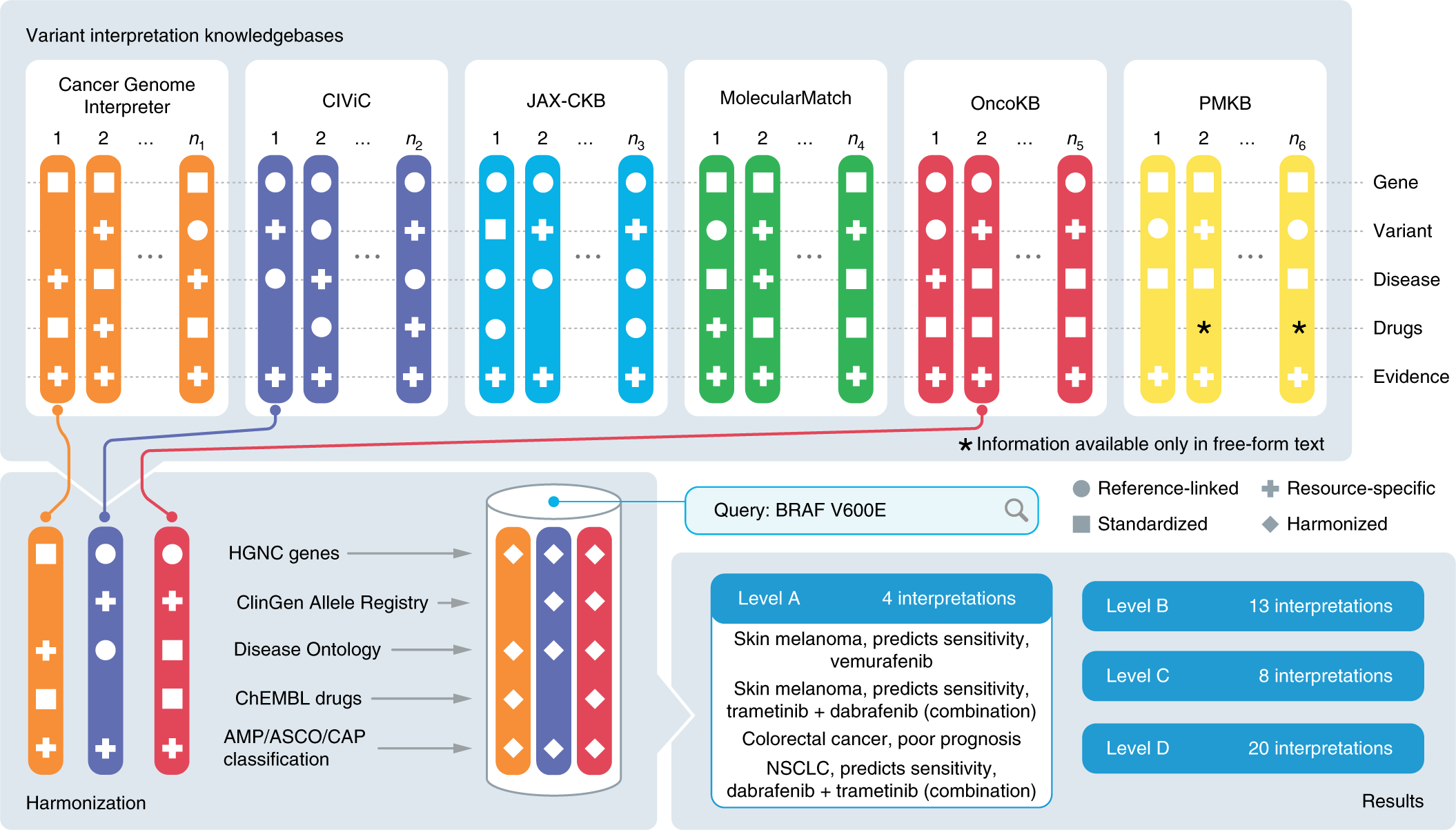
A harmonized meta-knowledgebase of clinical interpretations of somatic genomic variants in cancer | Nature Genetics
PLOS Computational Biology: GEMINI: Integrative Exploration of Genetic Variation and Genome Annotations
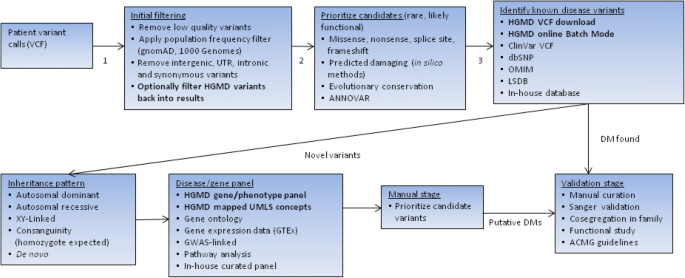
The Human Gene Mutation Database (HGMD®): optimizing its use in a clinical diagnostic or research setting | SpringerLink
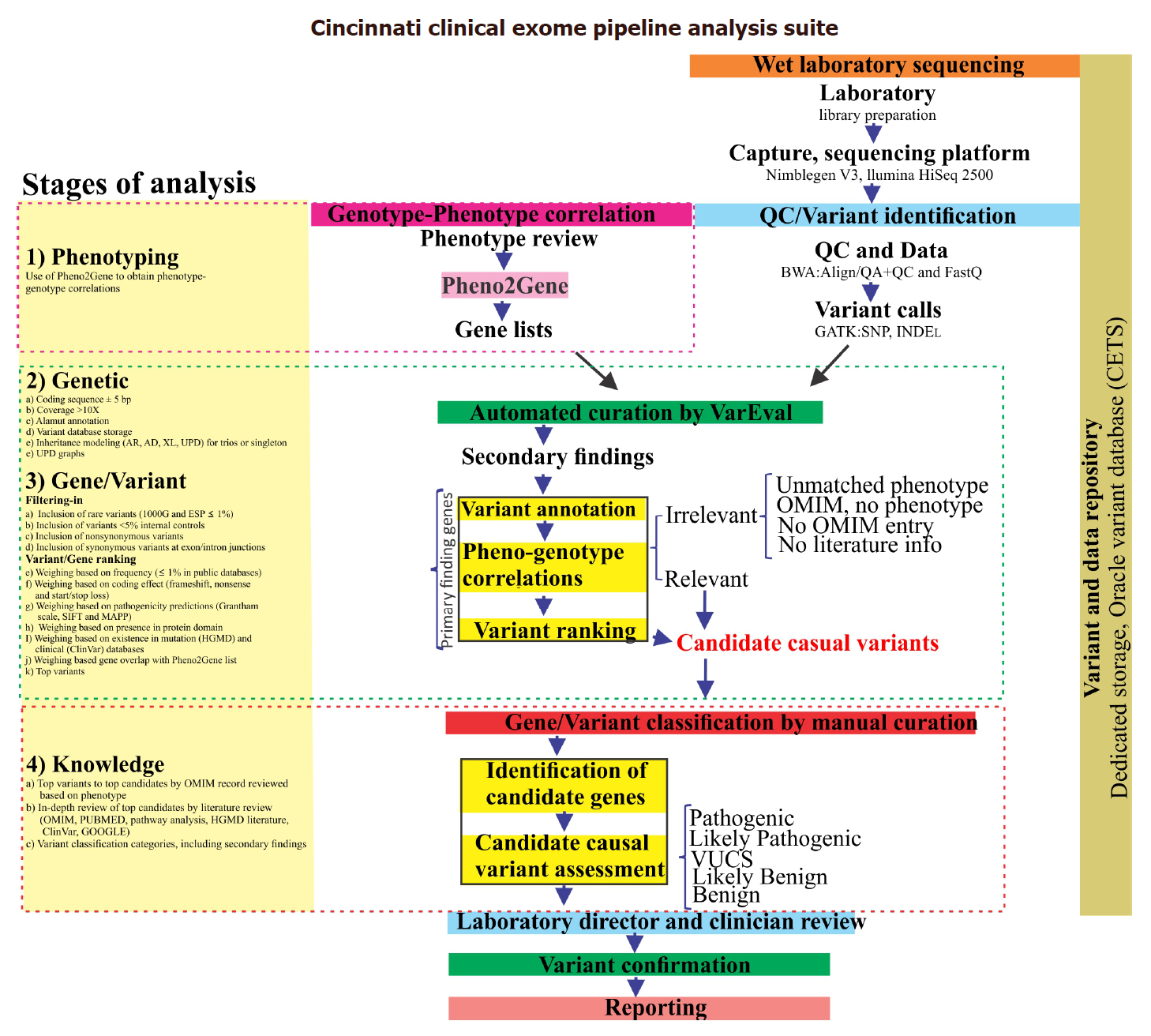
CCEPAS: the creation and validation of a fast and sensitive clinical whole exome analysis pipeline based on gene and variant ranking
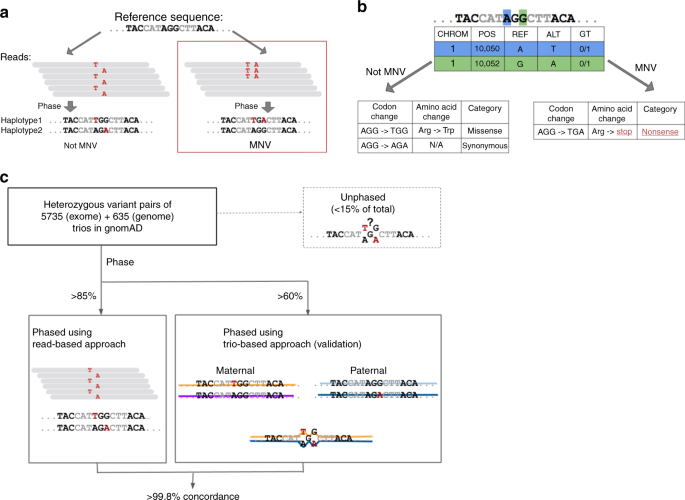
Landscape of multi-nucleotide variants in 125,748 human exomes and 15,708 genomes | Nature Communications

Technical desiderata for the integration of genomic data with clinical decision support - ScienceDirect
HuVarBase: A human variant database with comprehensive information at gene and protein levels | PLOS ONE

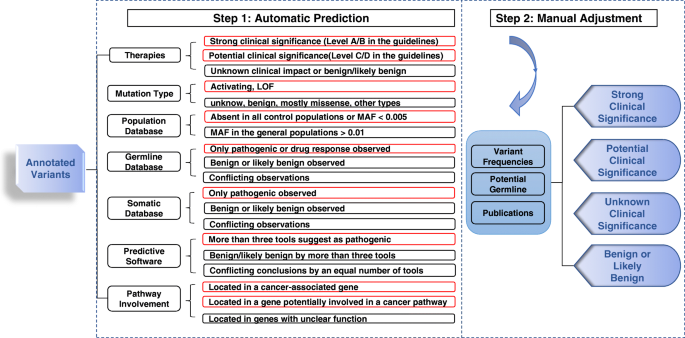
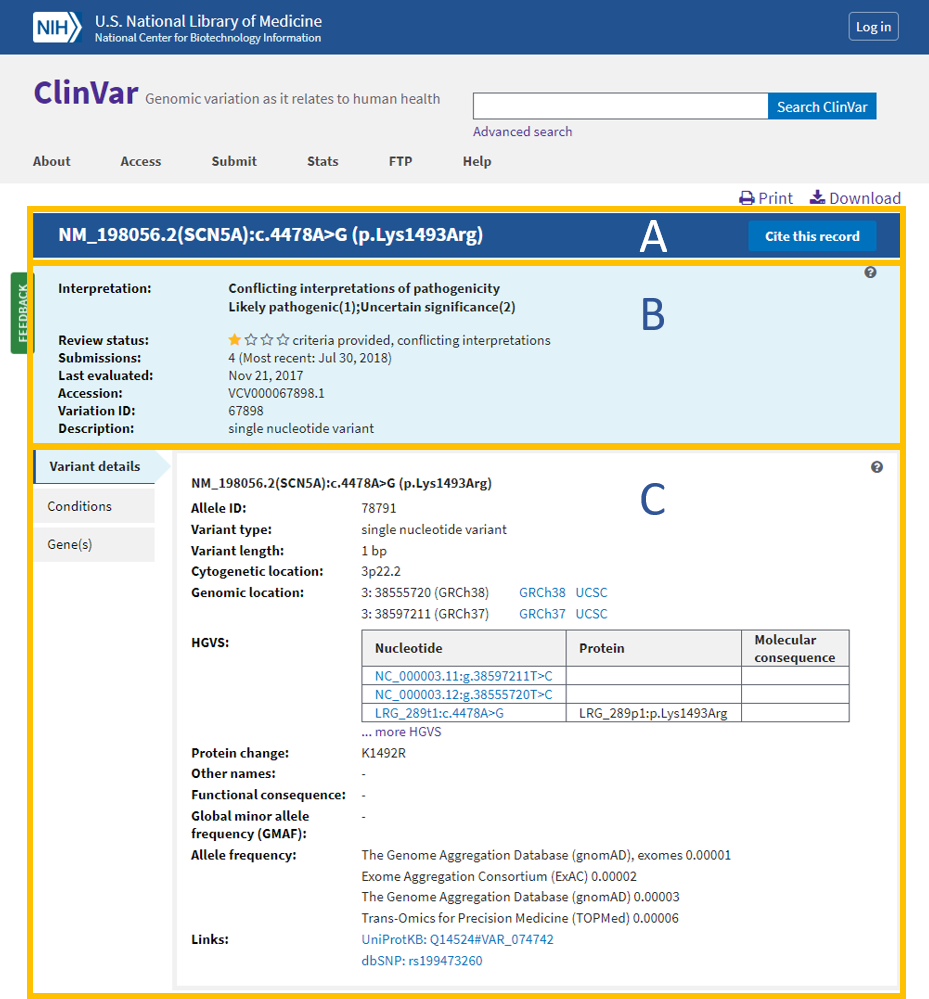
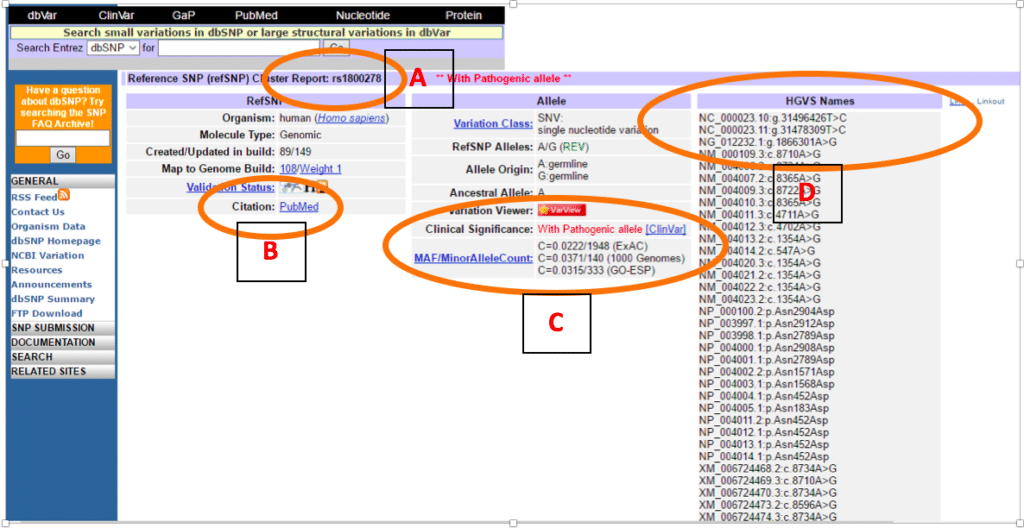
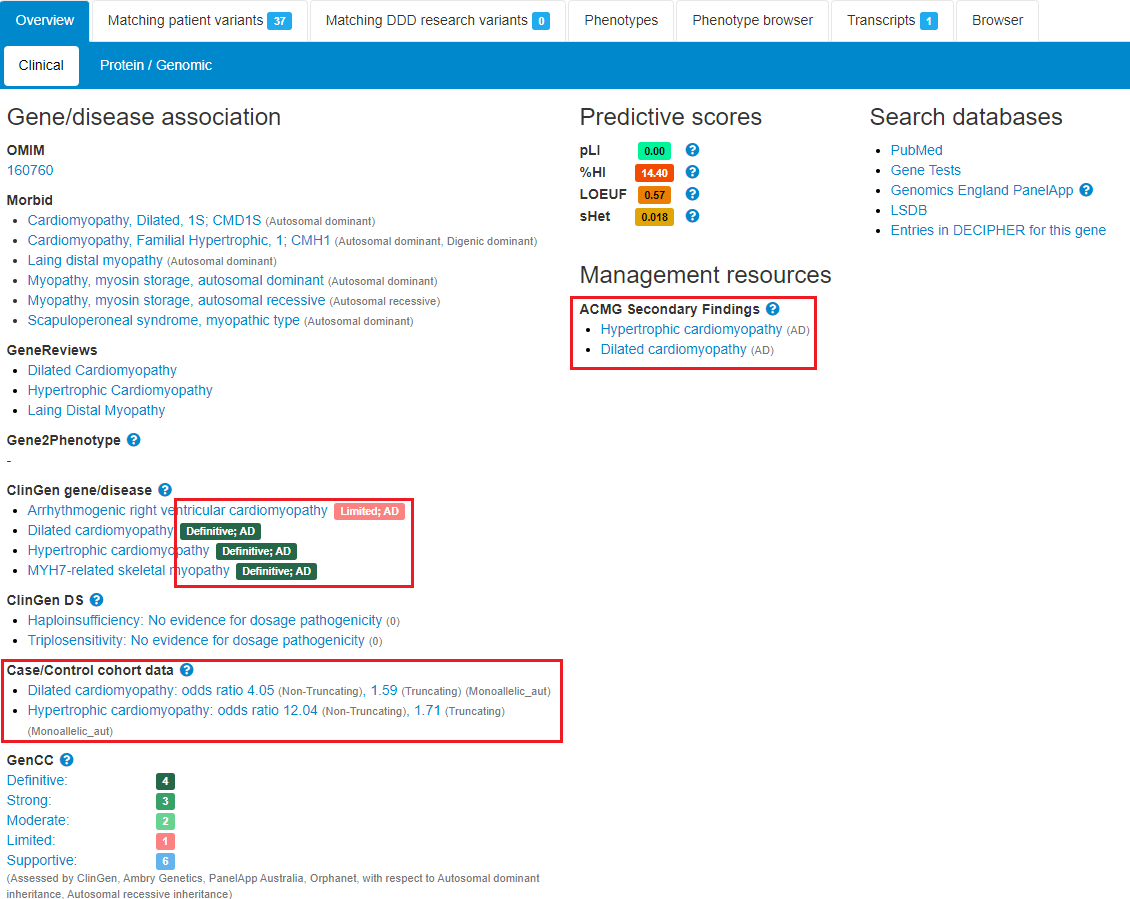
Comparison of Open-access Databases for Clinical Variant Interpretation in Cancer: A Case Study of MDS/AML | Cancer Genomics & Proteomics

Clinical Bioinformatics in Precise Diagnosis of Mitochondrial Disease - Clinics in Laboratory Medicine

Standards and Guidelines for the Interpretation and Reporting of Sequence Variants in Cancer - The Journal of Molecular Diagnostics